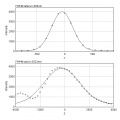
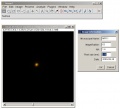
ASC Point Spread Function
(Created page with '**purpose: **requirements: **work flow: *overlay UV/V and VIS (confocal systems with separate UV/V laser coupling) **purpose: check the coupling precision of the UV/V laser **...') |
|||
| Line 1: | Line 1: | ||
| − | + | Important especially for high resolution objectives and deconvolution work. | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
== PSF Macro == | == PSF Macro == | ||
=== requirements for bead stacks === | === requirements for bead stacks === | ||
Revision as of 14:17, 30 September 2009
Important especially for high resolution objectives and deconvolution work.
Contents |
PSF Macro
requirements for bead stacks
- 100 planes, 200 nm spacing
- optimal: single bead in center of image
- the macro attempts to crop a 15x15 µm area with the bead in center; if the image dimensions are already smaller, extend the area: Fiji > Image > Adjust > Canvas Size...
install LUT
- Mac: go to applications folder > fiji: ctrl+click on the Fiji(.app) icon > Show package content
- PC: go to \Program Files\Fiji.app (or to the place you installed Fiji)
- create a folder called "luts", if it's not there yet
- drag and drop the *.lut into it
- (re)start fiji
install macro
- Fiji > Plugins > Macros > Install...
- select macro txt file
- macro gets installed to that menu for the current session
load bead stack
- Fiji > File > Open
- select bead stack file
- if necessary, split channels (Fiji > Image > Color > Split Channels)
- if necessary, crop image to a single bead
run macro
description by Laurent
Thanks a lot for your interest in our ImageJ Macro, which I join as an attachment.
You need to install it each time you start ImageJ. Go to >Plugins>Macros>Install...
Select it in the dialog window and click "open".
To run it, you need before to open a stack. We take stacks of 100 planes, spaced by 0.2mm, for all objectives and all microscopes. Then go to >Plugins>Macros>MIPs for PSFs for all microscopes to run the Macro.
Automatic Macro actions / User actions:
A. Selects the plane with the highest pixel intensity, adjusts display settings, opens the information dialog box.
1. Enter information in the dialog window which popped up.
2. Zoom in the image to clearly localize the center of the bead (you can also navigate between planes if needed).
3. Right clicks with the mouse on the center of the bead.
B. Crops the image to get 15mmx15mm area centered over the pixel clicked by the user.
C. Makes projections in X and Y of the stack
D. Stitches together the cropped area and the projections
E. Estimates and subtracts background
F. Takes the square root of the image (to minimize photon noise and to mimic a decrease in histogram gain)
G. Resizes the image to 550x550 pixels, adjusts display, changes LUT and saves the picture in a JPEG format with a standardized name: Date_Scope name_Magnification_NA.jpg.