Zeiss LSM 510 Tile Scan
Contents |
Overview
This manual demonstrates how to acquire Tile/Mosaic Scans with smooth transitions using a Zeiss LSM 510 confocal microscope. It uses the "Multi Time" macro.
Requirements
- Zeiss LSM 510 laser point scanning confocal microscope
- Scan Stage eg MCU28
- LSM Software v4.2
- Multi Time Macro 4.2 v1.2 (or higher...)
How To Do It
Using the "Stage" control
There is a tile scan option in the motorised stage controller window (Stage icon in the main LSM 510 software window). This works for single z planes only and has other limitations, but works in simple cases.
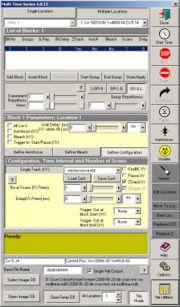
Using the Multi Time Macro
quick basic instructions
- Set up your imaging conditions. Define scan configuration and scan settings, adjust focal plane, signal intensity and image quality.
- Save your configuration in the according window, either single track or multi track.
- Make sure that you have a target database for the tile scan. If not, create a new database file via the main file menu.
- Open the multi time macro and select the multiple locations tab at the top.
- Select the first block (default setting) and click "define configuration". Choose single or multi track, select the saved configuration and load it via the "load config" button. Choose if you want to use the Z-stack feature by selecting the "Z-Stack" box. Stacks have to be set up as usual via the LSM software scan control.
- Define the tile scan area.
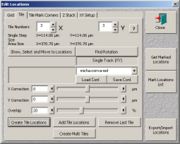
- (1. possibility) If you want to mark the center and arrange the tile scan around it (classic approach as in the stage utility), go to "edit locations" and select the "tile" tab. Drive the stage to the center of interest, define the tile numbers, set an overlap of 20% and click "create tile locations".
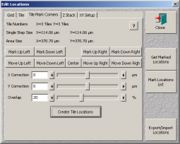
- (2. possibility) If you want to mark corners, go to "edit locations" and select the "tile mark corners" tab. Go to one corner with the scan stage and mark it using the according button. Drive the stage to the diagonally opposite corner and mark it. Set an overlap of 20% and click "create tile locations".
- Set the created database as the image database and set the "base file name". Select the "single file output" box.
- Start the tile scan with "start time". Wait for the acquisition and the automated saving.
- Select the remaining stack window "All_(base file name)_Sum".
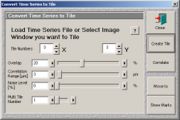
- Go to the multi time windows and click the "tile utilities". Set a correlation range of 3 µm.
- Click "correlate" and wait for the correlation. The processing time depends on the correlation range.
- Save the result. The "Save as" button in the image window disappears from time to time, in this case just save via the LSM software file menu.
Tile scanning instructions from Zeiss manual - might be outdated
copied out of an email to the cofocal list server on 22-02-2010 Julio Vazquez Fred Hutchinson Cancer Research Center Seattle, WA 98109-1024
Below is a summary of the procedures for collecting tiled images with
Zeiss LSM. I wrote this a long time ago, so there may be some errors,
or differences with different versions of LSM software, but it could
be a start.
Also, the LSM510 software acquires images by scanning a spot of light and reading intensities with a PMT, not a camera.
Zeiss LSM 510 META NLO User's Manual. Part III.
Advanced Techniques for multiple location imaging.
Contents:
I. Collecting a tile image. II. Calibrating stage. III. Marking Multiple locations. IV. Collecting Tiled Stacks. V. Collecting a grid of stacks. VI. Collecting stacks at multiple locations. VII. Collecting a Time series at multiple locations. VIII. More complex imaging patterns.
I. Collecting a tile image.
1. Set up your sample and imaging parameters. Optimize image.
2. In the main menu, open the Stage control panel: ACQUIRE>STAGE
3. At the bottom, under Tile Scan, enter the tile dimensions (n(x), n (y)). Maximum tile size is 4096x4096 pixels. Therefore, you can collect at most a 4x4 tile at 1024x1024 pixels, but 8x8 tile at 512x512. The lower the resolution, the larger the field of view you can collect.
4. Save your tile image.
Notes:
It is best to collect a small tile first at good resolution (e.g. 2x2 at 1024x1024 at zoom 1-2) to test stage alignment. If there is an offset in the tiles, proceed with stage calibration as described below.
This method only allows the collection in a single plane (not z
stacks). For tiled z stacks, see section IV.
II. Calibrating stage.
You need a sample that has good contrast.
1. Place sample on stage. Choose one channel that shows good image definition and brightness (overall good contrast).
2. Select objective and zoom factor. Optimize image.
3. In main menu, select ACQUIRE>STAGE, and set for tile collection under "Tile Scan", e.g. 2x2 tiles.
Collect an image and check alignment in x and y. If Alignment is good, the stage is calibrated: proceed with your experiment. Otherwise:
4. In the main menu, select MACROS>Rotation
5. In the calibration menu window, click "Calibrate". Microscope will start automatic scans and attempt to calibrate stage. When calibration is done, a correction factor will be displayed.
6. Collect a new tile to test the new calibration. Repeat if you are not happy with result. You may also try to change the numbers manually until satisfied. Note that alignment may not be perfect.
Note: Calibration will be best if you select "speed 1" in the Stage Position panel (Stage Control window).
III. Marking Multiple locations.
The Zeiss 510 allows you to mark locations in your sample and store
them. Once stored, you can navigate to any of those locations for
further imaging.
The recording of stage positions is done within the Stage control
window. You can mark positions in two different manners: manually,
or using a tile scan.
a. Manual marking:
1. You can select any position, for instance the center of your
sample, and mark that as the "center": Find the position you want to
be the center by visual inspection (e.g. you can use the 2.5x
objective to do a rapid survey of the sample.
2. In the stage window, under "Stage Position", click "Zero", and
click "Mark Pos."
3. Find a new location by visual inspection through the eyepiece, or
by moving the stage in fast scan mode. Find a new interesting
location. Click "Mark Pos.".
4. Repeat for all locations.
5. The list of locations will be recorded in the stage control
window, in the "Marks" pull-down menu.
6. You can now select a location in the pull-down menu, and use "Move To" to move the stage to that position.
Note:
The Stage control window also remembers the z position. Therefore, you can mark different locations in x,y,z. I noticed, however, that if you mark one position in x,y,z, the system will not record a new position that has the same x,y coordinates, but a different z location. Therefore, if you want to record two z positions at the same location, you will need to slightly move the stage in x or y.
b. Tile marking.
1. Collect a tile.
2. In the Tile Scan panel, click "Mark"
3. Click on your tile scan image to mark different locations.
Collection of complex images.
The Multi-Time Macro allows the collection of single z sections, or z
stacks, in a tiled manner, or on a grid pattern, or at multiple
locations. The various procedures are described below.
IV. Collecting Tiled Stacks.
1. Select a Multi-track configuration (not single track, as single
track do not remember certain settings).
2. Set your sample and optimize image collection. Adjust all parameters.
3. If you want z stacks, set the z stack parameters.
4. Save all the parameters as a new configuration: Click "Config", enter a new name (e.g. aa1), and "Store".
5. Calibrate stage as described in section II.
6. Start Multi-Time Macro (Main Menu>MACROS>MultiTime"R")
7. Click "Refresh" button.
8. Load Multi-Track configuration from pull-down menu in Multi-Time window (Config, Time-Interval and Bleach), (e.g. aa1)
9. For a single tile, make sure "number of scans" is set to "0", and z-stack is checked.
10. Under "List of Blocks", set number of repetitions to "0"
11. Make sure autofocus is NOT checked, and bleach is NOT checked.
12. Click "Single location" button on top of Multi-Time window.
13. Click "Edit Locations"
14. Select "Tile"
15. Enter the tile size (e.g. 2x2)
16. Select Overlap (e.g. 20%). The overlap is required for automatic stitching of images. If Stitching fails, use higher overlap.
17. Click "Create Tile"
18. Click "Options" (in Multi_Time window); Check "Keep Final Image Open" and "Save Final Image"
19. Click "Image Database", and select the database where your final image will be stored.
20. Click "Start Time"
21. Wait until images are collected and assembled. The final image will be saved in the database you selected. Intermediate images will be saved in a temporary directory in drive C.
V. Collecting a grid of stacks.
The "grid" function collects single sections or stacks in a grid pattern, with current position used as the origin.
To collect a grid, follow the same procedure as for a tile, except for:
14. Select "Grid"
15. Enter the spacing of the grid in x and y.
16. Click "Create Grid"
Proceed as for a tile with steps 18-21.
-- Julio Vazquez Fred Hutchinson Cancer Research Center Seattle, WA 98109-1024